Quite a difficult choice this year, with so many quality papers published. My favourite paper is actually a short series dealing with x-ray structure determination but with a difference! The key publication was submitted by Prof. Fujita of the University of Tokyo to Nature last year with a follow-up to Nature Protocols this year. The methodology basically means we can do away with the requirement of having perfect single crystals for x-ray structure determination and allows structure determination on as little as 80 ng of material.
From the authors” In our method, porous coordination networks are used as ‘crystalline sponges’ that absorb target molecules from the solution and orient them in a uniform fashion within the crystalline networks. Unlike many metal-organic frameworks, the pores of the crystalline sponges show a strong affinity for most organic molecules because of the sticky nature of the triazine ligand, namely, π-π , CH-π and charge-transfer interactions with guest molecules. The host-guest chemistry of the triazine ligand is well documented in discrete cage-host counterparts to infinitely networked crystalline sponges, and it facilitates the absorption of guest molecules from solution and their equilibrium placement in specific orientations in the pores of the coordination net-works. The newly ordered guest molecules within the single crystals of porous coordination networks can then be analyzed by X-ray analysis. We call these porous coordination networks that absorb and orient guest molecules for X-ray crystallography crystalline sponges, and we refer to our new crystal-free crystallography as the crystalline sponge method”.
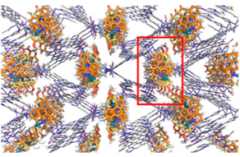
The crystalline sponge method still requires careful selection of the sponge crystal and one of the better was found to be
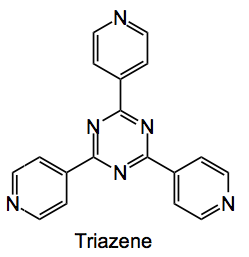
Which is a large metal-organic framework (MOF) formed by treating the triazene
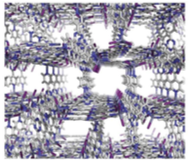
with zinc iodide in nitrobenzene methanol mixtures. Single crystals of the MOF grow on the glass of a test tube at the solution layer boundary and look like this:
Initially the pores are filled with nitrobenzene and this needs to be replaced with a non-coordinating solvent such as cyclohexane before the compound to be analysed can be introduced. This is done by simply soaking the MOF in cyclohexane with regular exchange for fresh solvent over the period of about a week. You obtain 3 types of crystal the best one to use is the rod-shaped crystal, which can be separated from the others by using a micropipette. They can be stored in cyclohexane for several months without degradation.
Now comes the time to add your guest compound. Using a saturated solution the guest is introduced to the MOF by slow evaporation favouring good diffusion into the pores . The progress of the diffusion is monitored by changes in the IR spectrum. Solvents are a bit limited here, cyclohexane being the optimal but small amounts of halogenated solvents can be employed as co-solvents. The micro-crystals can now be observed within the MOF unit cell by standard x-ray methodology. Recommended are synchrotron generated x-rays.
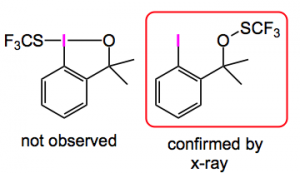
Since this was published I have seen three papers reporting the use of this methodology: one by Buchwald etal where it was used to reassign the structure of a hypervalent iodine reagent, for trifluorothiomethylation, from the benziodoxole structure to the open form shown below :
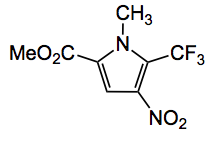
The other example I have seen is one from the Baran group, together with Prof. Fujita, where it was utilised to confirm the products of electrochemical C-H activation, for example the product of trifluoromethylation of the corresponding pyrrole:
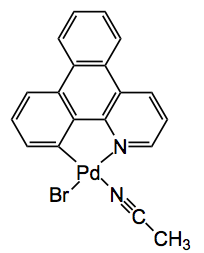
Finally the third application was one by Fujita himself where it was used to observe the palladium catalysed bromination of an aromatic compound with NBS taking place within the pores of the MOF! One of the intermediates seen was this compound, obtained after washing the RM with acetonitrile:
In a very elegant piece of work Fujita was able to follow the course of this reaction by x-ray crystallography. He observed a series of intermediates along the reaction pathway, in a time lapse sense. This sure beats TLC!!
While I was writing this a post appeared here. Apparently this method is being actively pursued to iron out the cracks as shown by this very recent paper. For example the procedure for the synthesis of the zinc based MOF used has been modified to yield crystals in 3 days instead of 2 weeks, quite a difference there. Other modifications are also in progress and presented in this paper.
Finally from Prof. Fujita “…. we successfully demonstrated the X-ray analysis of guaiazulene, a natural aromatic hydrocarbon, using an 80-ng sample and a standard laboratory X-ray machine. The nanogram-to- microgram scale X-ray measurement also enables researchers to implement a practical combination of HPLC separation and X-ray crystallographic analysis, known as liquid chromatography–SCD (LC-SCD). Furthermore, the absolute configuration of chiral compounds can be determined from the anomalous scattering effect of heavy atoms embedded in the framework of the crystalline sponges. Considering the importance of X-ray crystallography in structure analysis, the crystalline sponge technique is a major advance, and we expect it to be widely adopted in academic and industrial research.“
I agree with this statement entirely. Think about it, a single crystal x-ray of the compounds in your analytical HPLC separation. Couple this with the recently described MS/NMR methodology and you have a complete analysis of all the compounds produced by your reaction, including intermediates! This sponge x-ray method really advances the science of chemistry, not just organic but across the board. It is a wonderful tool and I hope it really takes off and becomes a part of everyday lab life.
Congratulations to Prof. Fujita and his group for providing this methodology.
![]()